Main content
Top content

Botany – Genetics and Evolution of Plant Diversity

Prof. Dr. rer. nat. Sabine Zachgo, Direktorin des Botanischen Gartens
Tel.: +49 541 969-2840
Fax: +49 541 969-2845
Sprechzeiten: n. V.
Raum: 35/E58
e-Mail
Research mission
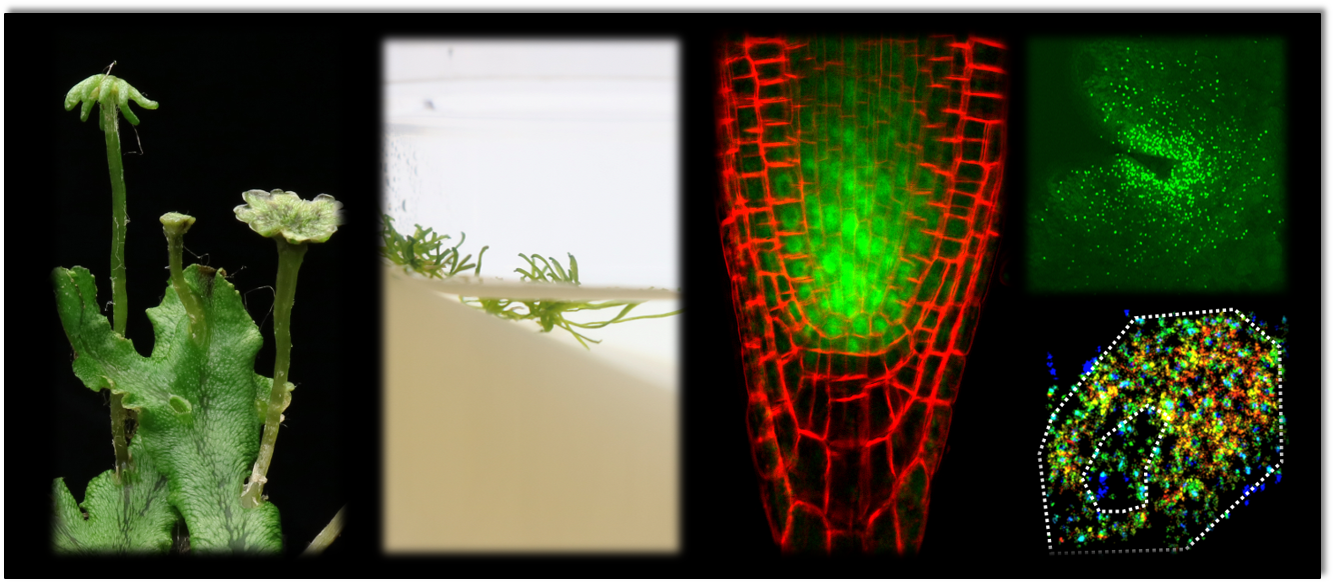
Our research team analyses key regulatory transcription factors that control plant development. We conduct comparative approaches using plant model organisms such as Arabidopsis thaliana together with other evolutionary informative plant species. Current research focuses on the liverwort Marchantia polymorpha and the amphibious liverwort Riccia fluitans to unravel the role of redox-regulation in developmental processes and for an adaptation to a terrestrial plant life style.
Model systems
- Arabidopsis thaliana and angiosperms with interesting novelties such as flower symmetry
- Marchantia polymorpha and Riccia fluitans
- Additional informative water and land plants for comparative studies
Methods
- Developmental genetics (genome editing, mutant analyses)
- Expression analyses (mRNA, protein)
- Transcriptomics (next generation sequencing)
- Morphological studies
- High resolution microscopy and redox sensors
- DNA/protein interactions (EMSAs
Selected publications
- Althoff F, Zachgo S (2020) Transformation of Riccia fluitans, an amphibious liverwort dynamically responding to environmental changes. International Journal of Molecular Sciences 21(15), 5410. DOI.org/10.3390/ijms21155410. pdf
- Maß L, Holtmannspötter M, Zachgo S (2020) Dual-color 3D-dSTORM colocalization and quantification of ROXY1 and RNAPII variants throughout the transcription cycle in root meristem nuclei. The Plant Journal, DOI 10.111/tpj.14986. pdf
- Busch A, Deckena M, Almeida-Trapp M, Kopischke, S, Kock, C, Schuessler, E, Tsiantis, M, Mithofer, A, Zachgo S (2019) MpTCP1 controls cell proliferation and redox processes in Marchantia polymorpha. New Phytologist, 224 (4), 1627-1641. DOI 10.1111/nph.16132. pdf
Further principal investigators
apl. Prof. Dr. Klaus Mummenhoff
apl. Prof. Dr. Barbara Neuffer